G4hunter Software Predicts G4 Rna Formation In The 3
We used the in-house G4Hunter algorithm to search for G4 putative sequences in a FASTA alignment of 107 HCV genomes RNA from the HCV genome database . A G4-prone fragment of the FASTA alignment was detected. This G4 sequence is located in the stem-loop IIy of the 3-end of the RNA . Interestingly, despite the high genetic variability of HCV, this sequence is highly conserved across 106 sequences from various HCV strains . The four blocks of Gs are always present, with only slight differences in length and base composition of the middle linker that separates the second and third G4 stretches. The sequence CGGGAGGGGGGGTCCTGGAGGC was the most frequently identified and occurred 65 times among the 106 sequences. In addition, 5 different motifs with G4Hunter scores ranging from 1.4 to 1.77 were found in the 42 other sequences . According to our experience with hundreds of sequences experimentally tested in our previous work, all of these sequences are very likely to form a stable G4 in vitro. Interestingly, Quadparser would have failed to pick them as two of the G-blocks contain only two consecutive guanines. This could explain why G4 formation in this region was overlooked using classical algorithms.
Table 1 G4 prone sequences detected by G4hunter. Mutations as compared to the most frequently found motif are shown in bold/italic.
Detection Of Extrahepatic Hcv Replication By Pcr
PCR is a very sensitive technique for detecting small amounts of nucleic acids. For detecting HCV, the viral RNA is converted to DNA by Reverse transcriptase , and then nested PCR is usually used to amplify particular regions of the genome. The two most commonly amplified regions are the 5’UTR and the hypervariable region 1 . The 5’UTR is a highly conserved region that is involved in viral replication and translation , while the HVR1 codes for the amino end of the envelope protein E2. E2 is involved in binding of HCV to host cells . Replication of HCV involves converting the viral genomic positive strand into an antigenomic negative strand, and then back to the genomic strand. Thus, the presence of the negative strand strongly suggests that replication is occurring.
We have only analyzed studies that reported RT-PCR of negative strands of HCV in humans. Many other studies have investigated positive strands, but the presence of positive strands is not an evidence of replication unless other data is presented that can only be found in cells replicating HCV. Some of those studies will be discussed in later sections. A number of studies have used model systems such as Chimpanzees and mice for analysis of HCV replication. We have not analyzed these studies, as they are model systems that have not proven useful for studying extrahepatic replication of HCV in humans.
Initial results with PCR
Hepatitis C Rna Pcr Qualitative Blood Test
The Hepatitis C RNA PCR Qualitative test is used to look for infections with the Hepatitis C virus. This test looks for the genetic material of the virus. Because viral genetic material may be detectable earlier than antibodies which develop in response to an infection, PCR testing can be used to screen for a recent exposure. This test is also useful as a confirmation for people who have had a positive result from a HCV Abs test. Results for this test are qualitative meaning they will come back as positive or negative.Hepatitis C is a virus spread through contact with infected blood. Nearly 80% of Hepatitis C infections develop into chronic Hepatitis. The number of people worldwide with chronic Hepatitis C infections is around 150 million. Chronic Hepatitis C infections can lead to serious health complication such as Cirrhosis and Liver Cancer. Many HCV infections display no symptoms. When symptoms do occur, some of the most common include:
Fever Grey feces Jaundice
Note: Result turn around times are an estimate and are not guaranteed. Our reference lab may need additional time due to weather, holidays, confirmation/repeat testing, or equipment maintenance.
Detection Period:
The Hepatitis C PCR test can typically detect the virus about 3 weeks from a suspected contact or exposure or anytime after. Some people may be detectable earlier.
Requirements:
Read Also: How Hepatitis C Affects The Body
Structure Of The Virus
The hepatitis B virion is a 42-nm particle comprising an electron-dense core 27 nm in diameter surrounded by an outer envelope of the surfaceprotein embedded in membranous lipid derived from the host cell . The surface antigen isproduced in excess by the infected hepatocytes and is secreted in the form of22-nm particles and tubular structures of the same diameter .
Interpretation of Results of Serologic Tests for Hepatitis B.
Hepatitis B surface antigen first appears during the late stages of theincubation period and is easily detectable by radioimmunoassay or enzymeimmunoassay. The antigen persists during the acute phase of the disease andsharply decreases when antibody to the surface antigen becomes detectable.Antibody of the IgM class to the core antigen is found in the serum after theonset of the clinical symptoms and slowly declines after recovery. Itspersistence at high titer suggests continuation of the infection. Core antibodyof the IgG class persists for many years and provides evidence of pastinfection.
Hcv Can Bind And Enter Extrahepatic Cells
The information below summarizes a number of studies that represents binding of selected viral proteins but they do not address the question of viral entry. One of the first studies on the subject of HCV in serum was on virus concentrated by ultracentrifugation, followed by testing the concentrate for positive and negative strands . These were tests done on cell free virus, and RNase and detergent sensitivity were tested. Positive strands were resistant to both RNase and detergents, while negative strands were resistant to either RNase or detergent. The investigators concluded that positive strands were probably protected in an enveloped core, while negative strands were membrane protected but not in a protein core.
Since much of the HCV in serum is complexed with antibodies, one study investigated infection of monocytic and T lymphocytic cell lines . They stimulated these cell lines with phorbal myristate and interferon- to stimulate Fc receptor expression before infecting with immune complexes of HCV. Entry was measured by RT-PCR. Non-stimulated cells showed no viral entry, while HCV negative strand was detected for up to 7 days after infection in monocytic cells. Little or no binding was seen for the lymphocytic cell lines. In the monocytic cell lines, monoclonal antibodies to FcRII, a low affinity immunoglobulin G receptor, abolished binding by immune complexes, suggesting that HCV can enter cells that express Fc receptors.
Recommended Reading: How Is Hepatitis B And C Transmitted
Protection Of Hepatitis B
The discovery of variation in the epitopes presented on the surface of thevirions and subviral particles identified several subtypes of HBV which differin their geographical distribution. All isolates of the virus share a commonepitope, a, which is a domain of the major surface proteinwhich is believed to protrude as a double loop from the surface of the particle.Two other pairs of mutually exclusive antigenic determinants, dor y and w or r, are alsopresent on the major surface protein. These variations have been correlated withsingle nucleotide changes in the surface ORF which lead to variation in singleamino acids in the protein. Four principal subtypes of HBV are recognized:adw, adr, ayw andayr. Subtype adw predominates in northernEurope, the Americas and Australasia and also is found in Africa and Asia.Subtype ayw is found in the Mediterranean region, easternEurope, northern and western Africa, the near East and the Indian subcontinent.In the Far East, adr predominates. But the rarerayr occasionally may be found in Japan and Papua NewGuinea.
Immunization against hepatitis B is now recognized as a high priority inpreventive medicine in all countries and strategies for immunization are beingrevised and universal vaccination of infants and adolescents is underexamination as a possible strategy to control the transmission of thisinfection. About 30 countries including the United States now offer hepatitis Bvaccine to all children.
Can I Take The Test At Home
At-home hepatitis C tests are available that allow patients to collect a blood sample at home and mail it to a laboratory for testing. Test samples are collected through pricking a finger with a sharp object, called a lancet, thats included in the test kit.
At-home HCV testing is a form of hepatitis C antibody testing and does not test for hepatitis C RNA or the strains genotype. Testing for hepatitis C at home is not a substitute for testing performed by a health care professional, and positive test results may need to be confirmed by laboratory-based testing.
Don’t Miss: Where Do I Get Hepatitis A Vaccine
Temporal Quasispecies Variation During Hcv Infection
Since different tissues contain different quasispecies, studies have investigated how these populations change over time. A recent study followed quasispecies found in four patients over a span of up to 18 years . They suggested there were four stages of HCV evolution: HCV establishes an infection incremental evolution of variants within subpopulations diversification into new subpopulations and strong negative selection in these subpopulations reduces variation and HCV achieves a stable adaptation to the host. Although the small number of patients in the study was a drawback, the model needs further investigation.
A different method of studying temporal variation is to study the repopulation of livers after transplantation. HCV from the cells in the serum, including monocytes/macrophages and lymphocytes, enter and populate transplanted liver tissue, and free virus in the blood also infects livers. We examined four studies that used RT-PCR to examine HCV infection after liver transplantation. Liver specimens had detectable amounts of negative strands by RT-PCR within 7 days after transplantation . Levels of negative strand in the liver after transplants do not correlate with serum levels of HCV , and negative strands were more likely to be found in transplanted PBMC than in PBMC from individuals with chronic HCV infection .
Signals In The Minus Strand
The secondary RNA structures that are conserved to be formed at the 5 end as well as at the 3 end of the antigenome minus strand usually do not represent exact mirror images of the structures formed by the plus strand . At the 5 end of the minus strand, the structure of the SL is well conserved, whereas the SL can assume three different structures . The biological relevance of these structures is not yet clear, and such possible relevance may be difficult to analyze since the counterpart of these sequences at the 3 end of the plus strand is extremely conserved and functionally relevant.
FIGURE 5. Elements in the 3 region of the HCV minus strand antigenome involved in plus strand synthesis. The black and gray lines and structures show the 3 region of the HCV minus strand antigenome. A contribution to plus strand initiation shown by in vitro-assays is indicated by boxes with solid green lines , a negative influence indicated by these in vitro-assays is indicated by a box with a red broken line . Sequences that were characterized by replicon studies to have a positive influence on RNA replication are boxed with dotted green lines . The region which is not boxed is variable in RNA secondary structure among the predictions , and no study showed a functional contribution of this region to replication. A possible relevance of the upstream structures SL VI and 588 as present in the minus strand is not clear.
Also Check: What Is The Treatment For Hepatitis
Distinctive Properties Of Hdv
The HDV particle is approximately 36 nm in diameter and composed of an RNA genomeassociated with HDAg, surrounded by an envelope of HBsAg. The HDV genome is aclosed circular RNA molecule of 1679 nucleotides and resembles those of thesatellite viroids and virusoids of plants and similarly seems to be replicatedby the host RNA polymerase II with autocatalytic cleavage and circularization ofthe progeny genomes via trans-esterification reactions. Consensus sequences of viroids which are believed to beinvolved in these processes also are conserved in HDV. Unlike the plant viroids,HDV codes for a protein, HDAg.
This is encoded in an open reading frame in the antigenomic RNA but four otheropen reading frames which are also present in the genome do not appear to beutilized. The antigen, which contains a nuclear localization signal, wasoriginally detected in the nuclei of infected hepatocytes and may be detected inserum only after stripping off the outer envelope of the virus withdetergent.
Hepatitis C Drug Targets Rna
An experimental drug developed by Danish startup Santaris effectively controls the hepatitis C virus in chimpanzees without creating drug-resistant forms of the virusa major advantage over other compounds in clinical development. The compound, a synthetic nucleic acid that binds to a microRNA molecule required for viral reproduction, is now in early-stage clinical trials. It is the first microRNA-targeting drug to be tested in humans.
Viral:
Approximately 170 million people across the globe are infected with the hepatitis C virus, a chronic infection that can lead to cirrhosis, liver cancer, and the need for a liver transplant. While drugs exist to treat the virus, they carry serious side effects and work in fewer than half of all infected patients. The treatment is very harsh and needs to be taken for 48 weeks, says Robert Lanford, the lead author on the new study, which was published online today in Science. Most people cant tolerate it that long, especially if they have liver disease.
Existing drugs suppress the virus by boosting the patients immune system. The Santaris drug targets the hepatitis C virus more directly by binding to a short piece of RNA called a microRNA, which the virus needs to replicate. The research is part of a larger effort over the last decade to develop methods of selectively targeting and silencing RNA molecules to treat a number of diseases.
Read Also: Ok Google How Do You Get Hepatitis C
Rnai Inhibits Hcv Protein Expression In Cultured Huh
We then examined the levels of HCV protein expression in siRNA-transfected S1179I cells by Western blot analysis. As shown in Fig. , levels of NS3 and NS5B proteins were unchanged at 2 days posttransfection , but they decreased on day 4 and day 6 posttransfection in S1179I cells transfected with NS3-1948 or NS5B-6133 compared with GAPDH-specific and scr siRNAs. At all time points analyzed, HCV protein expression in mock-transfected S1179I cells was not significantly different compared with S1179I cells transfected with the scr siRNA .
Effect of RNAi on IFN-induced genes. Real-time RT-PCR was used to compare the content of PKR and MxA transcripts in S1179I cells transfected with siRNA or incubated with IFN. The relative fold induction of each transcript normalized to that seen in mock-transfected S1179I cells is shown. Determination of expression of OAS and GAPDH mRNA by RT-PCR. Effect of IFN treatment and RNAi on HCV replication was assessed on day 2 after IFN treatment or siRNA transfection in S1179I cells, and compared with mock-transfected S1179I cells. Representative data from two independent experiments are shown.
Most Hcv Rna In Extracts Of Human Liver Is In Stable Dsrna Duplexes
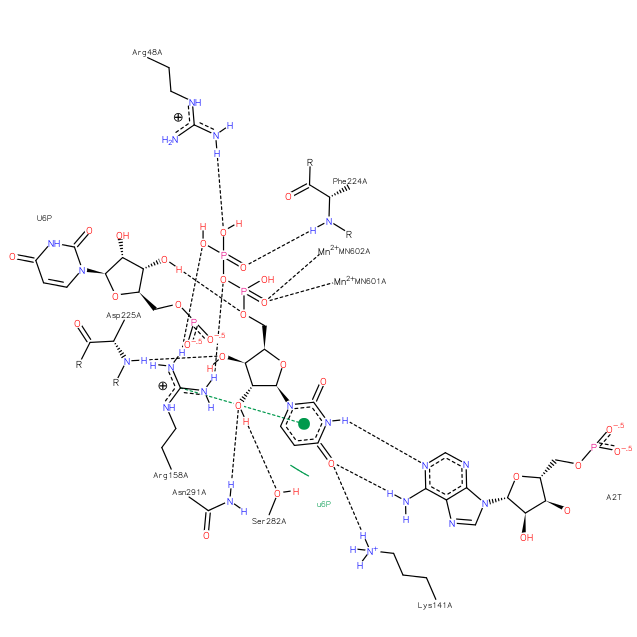
To enhance understanding of HCVhost interactions, we characterized HCV RNA in extracts of liver using techniques that fully separate the two strands of RNA duplexes before qRT/PCR. To achieve strand separation, RNA samples were heated to 106°C in 1 × TE and snapcooled in ice water to prevent reannealing. RNAs were then reversetranscribed under standard conditions. The resulting cDNAs represent the total population of HCV RNA in liver. This includes both the HCV RNA released from dsRNA as a result of heating to 106°C as well as the population of HCV RNA that was already in singlestranded form before heating to 106°C. For comparison, parallel samples were subjected to the standard RT reaction without having been heated to 106°C. These standard RT conditions were used to produce cDNA, which represents the population of free HCV RNA that was in singlestranded form before heating to 106°C.
Initially, extracts of human liver from HCV patients were analyzed using qRT/PCR targeting both the HCV plus and minus strand. Using an assay that detects the 5 untranslated region of the HCV plus strand and the complementary region of the minus strand , samples heated to 106°C had a 3.6fold higher signal than samples that did not undergo strand separation . Similar results were obtained using primers targeting the 3 UTR of the strand and the complementary region of the strand . Figure B summarizes the ability of various assays to detect free and duplexed forms of HCV RNA.
Don’t Miss: What Is Hepatic Luciferase Expression
Other Things To Know:
- The viral load measurement does not tell us anything about the severity of a patient’s liver disease or the degree of fibrosis . For that information, the patient would need additional testing.
- It is not necessary to check the viral load repeatedly during treatment.
- If a quantitative HCV RNA result is reported as “< 15 IU/L,” this means that the quantitative test cannot measure the hepatitis C virus. It may mean that there is no detectable HCV RNA at all, but it may mean that the level of virus is just too low for the test to pick it up.
Diagnosis Of Hcv Infection
Successful cloning of portions of the viral genome permitted the development ofnew diagnostic tests for infection by the virus. Since the original antigen wasdetected by antibodies in the serum of an infected patient it was an obviouscandidate for the basis of an ELISA to detect anti-HCV antibodies. A largerclone, C100, was assembled from a number of overlapping clones and expressed inyeast as a fusion protein using human superoxide dismutase sequences tofacilitate expression, and this fusion protein formed the basis of firstgeneration tests for HCV infection. The 5-1-1 antigen comprises amino acidsequences from the non-structural, NS4, region of the genome and C100 containsboth NS3 and NS4 sequences.
It is now known that antibodies to C100 are detected relatively late following anacute infection. Furthermore, the first generation ELISAs were associated with ahigh rate of false positive reactions when applied to low incidence populations,and there were further problems with some retrospective studies on stored sera.Data based on this test alone should, therefore, be interpreted withcaution.
The availability of the nucleotide sequence of HCV made possible the use of thepolymerase chain reaction as a direct test for the genome of the virus.There is considerable variation in nucleotide sequences among different isolatesof HCV, and the 5 non-coding region, which seems to be highlyconserved, is the preferred target for the PCR.
Read Also: How Is Hepatitis C Spread